Global profile mode
Sometimes all you want is a spatially-integrated spectrum of a source. The
GlobalProfile class offers a simplified setup of MARTINI for
this purpose.
Assumptions and limitations
In order to offer a simpler and faster way to produce a spectrum, the
GlobalProfile class makes some assumptions:
No spatial aperture is assumed. Every particle in the source contributes to the spectrum (unless it falls outside of the spectral bandwidth).
The positions of particles are still used to calculate the line-of-sight vector and the velocity along this direction.
There is therefore no need or way to specify a beam or SPH kernel as with the main
Martini class. It is also not possible to use MARTINI’s noise
modules with this class. If these restrictions are found to be too limiting, the best
course of action is to produce a spatially-resolved mock observation and derive the
spectrum from those data as would be done with “real” observations. For example, if the
spectrum within a spatial mask defined by a signal-to-noise or other cut is desired, or if
spatially-dependent effects like primary beam attenuation are relevant, then the
GlobalProfile class should not be used. The
GlobalProfile class is mainly intended to efficiently provide a
“quick look” at the spectrum, or a reference “ideal” spectrum.
Usage
The source and spectral model
modules should be set up as for a full MARTINI mock observation. The
beam, noise and
sph_kernel modules are not relevant. The
datacube module is not used, but a subset of its configuration
options are instead given directly to the GlobalProfile.
Schematically, an example initialisation looks like:
from martini import GlobalProfile
from martini.sources import SPHSource
from martini.spectral_models import GaussianSpectrum
source = SPHSource(...)
spectral_model = GaussianSpectrum(...)
gp = GlobalProfile(
source=source,
spectral_model=spectral_model,
n_channels=64,
channel_width=10 * U.km * U.s**-1,
spectral_centre=source.vsys,
)
The arguments to the other modules are omitted here (replaced with ...), check the
documentation pages of each module for details. Here the spectrum will be centred on the
source systemic velocity, but an explicit frequency or Doppler velocity value could be
given instead. The units of the channel_width argument determines whether the
resulting spectrum will have channels evenly spaced in frequency (if channel_width has
dimensions of frequency) or velocity (if it has dimensions of speed).
Inserting the source
The spectrum can be accessed through the spectrum attribute:
gp.spectrum
If it has not yet been calculated, it will be calculated when accessed. The calculation can be explicitly forced with:
gp.insert_source_in_spectrum()
but this is not usually necessary. In addition to the spectrum itself, the centres and edges of the channels are available as:
gp.channel_mids
gp.channel_edges
respectively. These arrays will have dimensions of frequency or velocity depending on the
units of the channel_width argument passed to GlobalProfile
on initialization. You can obtain the channel edges or centres in your preferred
dimensions with:
gp.frequency_channel_mids
gp.frequency_channel_edges
gp.velocity_channel_mids
gp.velocity_channel_edges
Parallelization
The core loop in the source insertion function is a loop over pixels. Since
parallelization is implemented for this loop, and for a
GlobalProfile there is a single pixel, parallelization is not
available in this mode.
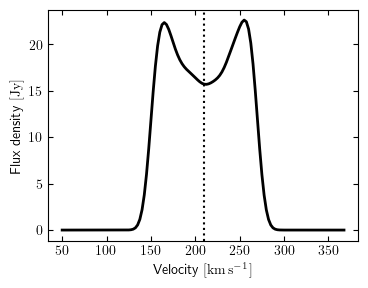
Quick plot of the spectrum
As a convenience, a function is provided to make a quick plot showing the spectrum.
Whether the systemic velocity of the source (as reported by source.vsys) is shown by a
vertical dotted line is controlled by the show_vsys argument. The channels
argument (either "velocity" or "frequency") determines the units of the spectral
axis in the figure.
from martini import demo_source, GlobalProfile
from martini.spectral_models import GaussianSpectrum
source = demo_source(N=20000) # create a simple disc with 20000 particles
gp = GlobalProfile(
source=source,
spectral_model=GaussianSpectrum(sigma=7 * U.km * U.s**-1),
n_channels=128,
channel_width=2.5 * U.km * U.s**-1,
spectral_centre=source.vsys,
channels="velocity",
)
gp.plot_spectrum(
show_vsys=True,
)
The resulting figure is returned by the function, or can be directly saved to file with
the save argument (e.g. save="myplot.png" or save="myplot.pdf"). This example
looks like: